Pattern recognition receptors (PRRs) are core immune sentinels in plants that defend against pathogens. Among them, the leucine-rich repeat receptor-like kinase subfamily XII (LRR-RLK-XII) has become a key research target in plant genetic improvement due to its strong species specificity and great potential for disease resistance across different crops. However, traditional genetic tools can only identify PRRs in model plants such as Arabidopsis thaliana. In non-model plants (such as perennial crops and forest trees), PRR discovery is severely hampered by difficulties in genetic transformation, long generation cycles, and incomplete genomic information. Simultaneously, existing PRRs suffer from narrow ligand recognition ranges and low modification efficiency, making it difficult to cope with constantly mutating pathogens and failing to meet the disease resistance needs of agricultural production.
On September 4, 2025, a team led by Ken Shirasu from the University of Tokyo and Yasuhiro Kadota from the RIKEN Center for Sustainable Resource Science published a research paper in Science titled "Systematic discovery and engineering of synthetic immune receptors in plants." This study integrated bioinformatics, synthetic biology, and biochemistry techniques to construct a large-scale LRR-RLK-XII receptor screening and engineering system. They successfully identified a novel cold shock protein receptor, SCORE, and through structural analysis and site-directed mutagenesis, obtained synthetic receptor variants capable of recognizing multiple crop pathogens. This work represents the first successful efficient discovery and precise engineering of immune receptors in non-model plants, providing a new solution for disease-resistant breeding in perennial crops.
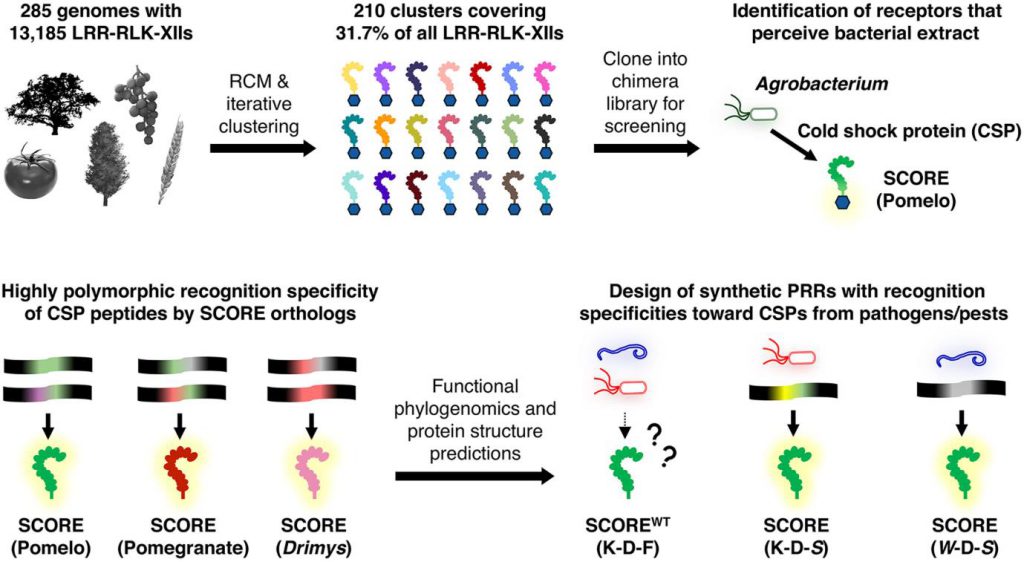
Figure 1. Pipeline for systematic discovery and engineering of plant immune receptors. (Ngou, et al. 2025)
First, the authors collected 13,185 LRR-RLK-XII receptors from the genomes of 285 angiosperm species, focusing on the species-specific expansion characteristics of the LRR-RLK-XII family. Through repetitive sequence conservation mapping (RCM) analysis, they focused on non-conserved residues in the extracellular domain that determine ligand recognition specificity. After multiple rounds of clustering and screening, they obtained 210 highly conserved subgroups (covering 31.7% of the target receptors), significantly narrowing the scope of functional verification. To avoid interference from endogenous plant immune responses, they designed LRR ectodomain-BRI1 kinase domain chimeric receptors. When the receptor transiently expressed in Nicotiana benthamiana recognizes a pathogen ligand, it triggers BES1 protein dephosphorylation signaling through the BRI1 kinase domain, which can be directly detected by Western blot without relying on complex immune signaling analysis. Using this system, they screened the chimeric receptors from the 210 subgroups and ultimately discovered 7 positive receptors that could be activated by Agrobacterium. Among them, receptor 181 from grapefruit showed the best performance, recognizing heat extracts from various bacteria, becoming the core subject of subsequent research.
Through protein isolation, anion exchange chromatography, size exclusion chromatography, and mass spectrometry analysis, the authors discovered that receptor 181 specifically recognizes a highly conserved 15-amino acid peptide (csp15) in bacterial cold shock proteins (CSPs). Although this receptor is functionally similar to the previously identified Solanaceae-specific CSP receptor CORE, their evolutionary paths are completely independent. The intracellular LRR surface consistency between the two is only 18.8%, and there are significant differences in their recognition mechanisms (CORE is sensitive to C-terminal truncation, while SCORE is sensitive to N-terminal truncation). Therefore, it was named Selective Cold Shock Protein Receptor (SCORE). Further research showed that SCORE and its homologs are mainly distributed in Sapindales (such as citrus), Malvales (such as cotton), and Magnoliales plants, showing clear lineage specificity. The authors cloned 20 SCORE homologs from more than 60 plant species, of which 13 stably recognized different csp15 variants (including SCORE from pomelo, i.e., CM), 4 showed weak recognition, and 3 were inactive. This functional polymorphism stems from differences in the 6th and 7th amino acid residues of csp15 and is a direct result of natural selection in plants responding to different pathogens.
Based on AlphaFold2 structure prediction, the authors clarified the interaction mechanism between SCORE (CM) and csp15: the 3rd to 16th repeat units of the leucine-rich repeat (LRR) domain of SCORE are involved in ligand binding. The N-terminus (valine, lysine) binds to glycine and charged/aromatic residues of the 13th-16th LRR, the C-terminal GFGF motif interacts with the conserved QSEMQSY segment of the 6th-7th LRR, while the 6th-7th variable residues of csp15 (e.g., Ala/Glu) form key specificity sites with the 8th-11th LRR residues. The diversity of the 8th-11th LRR residues (especially positions 300, 302, and 304 of the 10th LRR) in different SCORE paralogs leads to differences in ligand recognition. Experimental validation showed that constructing chimeras by exchanging LRR motifs revealed that introducing the 10th LRR motif from different homologs into CM significantly altered its ligand recognition spectrum, confirming that this motif is a switch for SCORE specificity.
By analyzing the surface charge distribution of SCORE paralogs, the study found that the inner surface charge characteristics of the 8th-11th LRR motifs (determined by residues 300, 302, and 304) directly regulate csp15 recognition specificity. For example, SCORE that preferentially recognizes negatively charged csp15 (SCORE from pomegranate, PG) exhibits a positively charged surface, while receptors that recognize neutral csp15 (such as CM) have a neutral or mixed charge. By site-directed mutagenesis of these key residues in SCORE, its charge properties can be systematically altered, successfully constructing 37 synthetic variants. Experiments showed that wild-type CM could not recognize CSPs from pathogens such as Ralstonia solanacearum, citrus greening disease pathogen, and root-knot nematodes, while variants with 1-4 amino acid mutations could efficiently recognize the csp15 peptides of these pathogens. For example, a single residue substitution significantly increased CM's sensitivity to DE-type csp15. This study demonstrates that by targeting and reshaping the charge of key sites on the inner surface of LRR, synthetic immune receptors with broad-spectrum resistance to diseases and pests can be precisely engineered.
The core innovation of this study lies in the construction of a complete technological system encompassing whole-genome clustering, chimeric receptor screening, structural analysis, and site-directed mutagenesis, enabling the large-scale discovery and precise engineering of immune receptors in non-model plants for the first time. This system overcomes the limitations of traditional genetic tools, allowing for the rapid identification of functional PRRs without relying on genetic transformation of non-model plants. The discovery and modification of SCORE receptors not only reveal the evolutionary diversity and ligand recognition mechanisms of plant PRRs but also provide novel disease resistance tools for perennial crop diseases such as citrus greening disease. Furthermore, the study demonstrates that the ligand specificity of PRRs can be altered through mutations in a few key residues, providing a clear paradigm for the targeted design of plant immune receptors. Its modular design approach can also be extended to other PRR families, opening new avenues for broad-spectrum disease resistance breeding in crops.