Lifeasible specializes in RNA-protein interaction study. We offer several professional and advanced RNA-protein interaction analytical technologies. Of which, protein-RNA interaction mapping assay (PRIMA) is a translation-based assay that examines protein-RNA interactions in the yeast cytoplasm. PRIMA allows the pairwise testing of an RNA “bait” versus multiple RBP “preys” in a single experiment.
The principle of PRIMA depends on that yeast poly(A)-binding protein (Pab1p) can bind the 3’ poly(A) and interact with the 5’ end of an mRNA with the help of the scaffold protein, eIF4G, and the cap binding protein, eIF4E. These interactions can stabilize the mRNA and increase translation of the mRNA into protein. In PRIMA, a reporter mRNA that encodes GFP is used and its poly (A) tail is replaced with a selected RNA “bait” element of interest. RBP “preys” are fused to Pab1p. When the RBP binds the RNA “bait” element, Pab1p interacts with the 5’ end of the reporter mRNA through eIF4G and eIF4E, resulting in stabilization and production of GFP (Figure 1A). The GFP fluorescent signal can be measured in a quantitative manner using high-throughput flow cytometry, and the strengths of positive interactions are calculated using computational data processing and statistical analyses of replicates (Figure 1B).
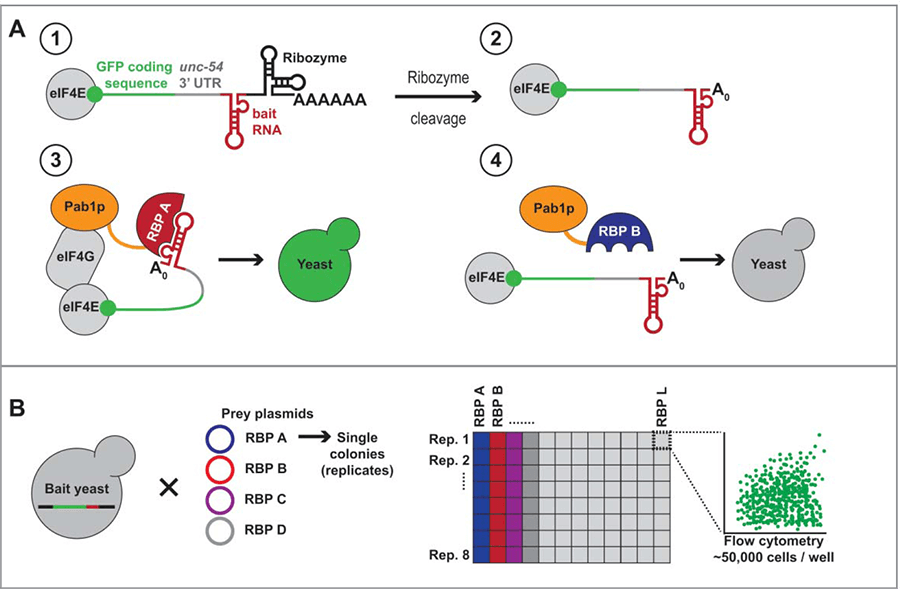 Figure 1. PRIMA design and experimental workflow (Tamburino et al., 2017).
Figure 1. PRIMA design and experimental workflow (Tamburino et al., 2017).
With years of experience and excellent staff in PRIMA, Lifeasible has developed a unique expertise of handling all steps involved from vector construction, yeast transformation to data collection and analysis. Lifeasible always works together with customers to solve specific challenges and chase highest-level customer satisfaction. Welcome to contact us for more details.
Reference