Professional, Customizable Medicago truncatula Transformation Services
Lifeasible delivers specialized genetic transformation solutions for Medicago truncatula (Barrel Medic), a cornerstone model organism for legume biology and symbiosis research. With its compact diploid genome (approximately 450–500 Mb), short generation time (about 3 months), and exceptional capacity for nitrogen-fixing nodule formation, M. truncatula serves as the premier system for dissecting plant-microbe interactions and root developmental biology. Our end-to-end platform bridges the gap between vector design and stable transgenic lines, empowering researchers to explore rhizobial symbiosis, root architecture, and specialized metabolite pathways with unprecedented efficiency.
Leveraging our deep expertise in plant genetic engineering, we support diverse project objectives—from CRISPR-based knockout of symbiosis-related genes to heterologous expression of pharmaceutical proteins in root cultures.
TARGET GENOTYPES
R108-1, Jemalong A17, etc.
Standard ecotypes and regenerable variants available
TYPICAL YIELD
8–25
Independent T0 Positive Events per construct
EDITING EFFICIENCY
Up to 75%
High-efficiency CRISPR/Cas9-mediated gene targeting
LEAD TIME
5–7 Months
From vector receipt to T1 seed collection
Standard Package
Efficiency Focused
Premium Package
Full-Service Custody
Stable transformation constitutes the foundation of modern legume functional genomics, enabling heritable integration of novel traits into the M. truncatula germline. At Lifeasible, we have refined Agrobacterium-mediated protocols to accommodate the unique regeneration requirements of Barrel Medic, ensuring robust T-DNA integration with minimal copy number complexity.
While our primary methodology relies on Agrobacterium tumefaciens for precise genomic insertion, we also provide specialized alternatives for challenging genotypes or large DNA constructs exceeding conventional T-DNA limits, including protoplast-based transformation for transient validation prior to stable line commitment.
![]()
Explant Preparation
Prepare cotyledon nodes, leaf discs, or embryogenic suspensions.
![]()
Infection & Co-cultivation
Infect explants with Agrobacterium and co-cultivate 2–5 days.
![]()
Stringent Selection
Select transformants using Kanamycin, Hygromycin, or Basta antibiotics.
![]()
Morphogenesis & Regeneration
Regenerate shoots and roots via hormone optimization.
![]()
Acclimatization & Phenotyping
Acclimatize plantlets and test nodulation competence.
For projects requiring rapid functional validation without the extensive timeline of stable transformation, Lifeasible provides high-efficiency transient expression systems for M. truncatula. These assays enable quick assessment of gene constructs, promoter strength analysis, protein subcellular localization, and CRISPR guide RNA efficiency within days rather than months.
![]()
Vector Design & Preparation
Select vectors and prepare high-purity plasmids.
![]()
Target Material Isolation
Isolate protoplasts or cell suspensions from leaves.
![]()
DNA Delivery
Transfect cells using PEG or electroporation
![]()
Incubation & Analysis
Incubate cells and analyze via imaging or blotting.
Lifeasible deploys refined methodologies addressing the unique cell wall composition and regeneration requirements of Medicago species.
Utilizing disarmed Agrobacterium tumefaciens strains (EHA105, GV3101, AGL1) with acetosyringone induction, we achieve high-efficiency T-DNA transfer into cotyledon-derived callus. Vacuum infiltration alternatives for cell suspension cultures enable high-throughput screening without solid medium requirements.
PEG-mediated transfection of mesophyll protoplasts or electroporation of cell suspension cultures for rapid, DNA-free validation assays. Ideal for CRISPR efficiency screening and protein localization studies.
| Category | Requirements |
| Sample Type | Surface-sterilized mature seeds, sterile explants, or established cell suspension cultures |
| Sample Amount | Minimum 100–200 healthy seeds (approximately 3–5 g) or 50–100 explant pieces |
| Pre-Treatment | Scarification recommended for hard-seeded lines; seeds must be free from fungal contamination and chemical treatments |
| Storage Conditions | 4°C dry storage for seeds; avoid long-term storage (>12 months) for lines with regeneration requirements |
| Shipping | Ambient temperature with desiccant for seeds; active cultures require insulated cold packs |
| Metadata Needed | Accession/ecotype name, generation status, known transformation recalcitrance, target gene/construct details, preferred selection markers |
| Vector Information | Complete plasmid maps including promoters (CaMV35s, legume-specific, or inducible), GOI, selection cassettes, and reporter configurations |
Enhance your core transformation project with specialized downstream capabilities:
Molecular Characterization & Validation
Southern blotting for insertion copy number, RT-qPCR for transcript quantification, and subcellular localization using GFP/RFP fusions.
CRISPR/Cas9 Off-Target Analysis
Whole-genome sequencing or targeted amplicon sequencing to verify editing precision and identify potential off-target modifications.
Custom Vector Engineering
Design of tissue-specific expression cassettes (root-specific promoters, nodule-specific elements), multi-gene stacking vectors, and CRISPR/Cas12a systems for multiplexed editing.
Symbiosis-Specific Assays
Quantitative nodulation kinetics, nitrogen fixation efficiency measurements (acetylene reduction), and mycorrhizal colonization studies for arbuscular symbiosis research.
Hairy Root Culture Development
Agrobacterium rhizogenes-mediated transformation for rapid root biomass production and compound screening without whole-plant regeneration.
Vector Construction & Optimization
Explant Preparation & Pre-culture
Transformation & Selection
Regeneration & Hardening
Molecular Characterization
Seed Production & Harvest
Note: Timelines vary based on genotype regeneration speed (R108-1 typically faster than Jemalong) and project complexity (CRISPR projects may require additional screening).
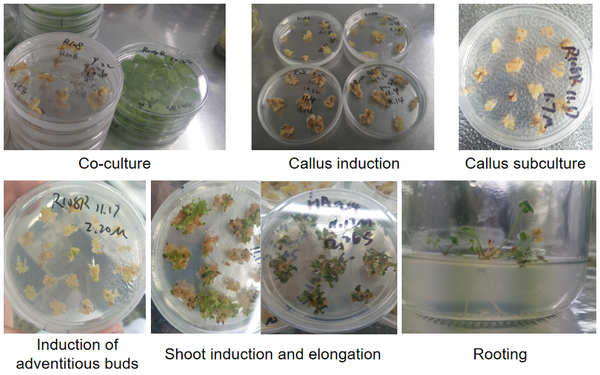

Standard Stable Transformation Pipeline
This internal project demonstrates the complete regeneration trajectory of Medicago truncatula R108-1 via somatic embryogenesis following Agrobacterium-mediated transformation. The visual timeline captures the critical developmental stages: co-cultivation of cotyledon explants with GV3101 harboring a binary vector, subsequent callus induction on selective medium, and iterative subculture to establish embryogenic cultures. The progression continues with induction of adventitious buds, shoot elongation under hormone-optimized conditions, and finally robust rooting to generate soil-ready T0 plantlets. This standardized workflow consistently yields 15–20 independent transgenic events per construct, with typical transformation efficiency of 60–75% for this highly regenerable ecotype.

Streamlined Agrobacterium-Mediated Transformation of M. truncatula A17
This suspension culture-based protocol eliminates specialized equipment requirements, delivering stable transgenic lines in approximately 12 weeks without vacuum infiltration or protoplast preparation. By selecting day-7 cells (early exponential phase) and utilizing a simplified 3-day co-cultivation workflow, the method reduces contamination risks and technical barriers associated with conventional callus-based approaches. Validation using SARS-CoV-2 Spike protein RBD demonstrated consistent expression (~1.6 mg/L) with lower variability than traditional methods, confirmed through PCR and Western blotting. This accessible platform enables rapid recombinant protein production and functional screening in legume cell cultures without advanced tissue culture infrastructure.
Our commitment to precision and reliability has established Lifeasible as a trusted partner for academic and industrial legume researchers globally:
"Lifeasible generated 20 independent T0 lines for our R108-1 CRISPR project within 6 months. Critically, 16 lines retained full nodulation capacity with S. meliloti, allowing immediate assessment of root infection phenotypes. The cotyledon-based transformation approach delivered exceptional efficiency."
Dr. A. Schmidt
Principal Investigator
USA
"We needed rapid validation of seven different promoter constructs driving root-specific expression. The transient protoplast system provided clear fluorescence data within 72 hours, saving us months of stable transformation work. Data quality matched our in planta observations perfectly."
Dr. J. Tanaka
Group Leader
USA
"Transitioning from Arabidopsis to Medicago was seamless with Lifeasible's guidance. The Premium Package Southern blot analysis confirmed single-copy insertions in 9/12 A17 lines, and the nodulation assays confirmed that tissue culture didn't compromise symbiotic signaling."
Dr. R. Moreno
Research Fellow
UK
Legume Specialization
Deep technical expertise in Medicago biology, including ecotype-specific regeneration requirements and preservation of symbiotic competence through tissue culture.
Genotype Versatility
Proven transformation success across major reference lines (R108-1, Jemalong A17/J5, Jester) and specialized variants, with customized protocols for each genetic background.
Integrated Symbiosis Support
Unique capability to validate transgenic lines through rhizobial inoculation and nodulation assays, providing functional confirmation beyond simple transgene detection.
Regulatory Compliance
All operations conducted within certified biosafety facilities, ensuring containment standards appropriate for transgenic legume research and international shipping requirements.
Ready to accelerate your legume research program?
Our technical specialists are prepared to discuss your specific project requirements, from single-gene knockouts to complex metabolic engineering in Medicago truncatula. Whether investigating the molecular basis of nitrogen fixation or developing novel biopharmaceutical production platforms, Lifeasible provides comprehensive support from construct design to phenotypic validation.
Medicago truncatula serves as the definitive bridge between model plant systems and agriculturally critical legume crops:
Early Medicago genetic engineering historically confronted significant genotype-specific regeneration barriers, with many ecotypes exhibiting recalcitrant tissue culture responses and unpredictable somaclonal variation that limited experimental throughput. Contemporary protocols strategically leverage the highly regenerable R108-1 ecotype for routine high-throughput applications while simultaneously optimizing co-cultivation conditions and hormone balances for the sequenced Jemalong A17 reference standard. Recent technical innovations have dramatically expanded transformation capabilities, including cell suspension-based rapid screening platforms for accelerated validation, protoplast regeneration systems enabling DNA-free genome editing approaches, and in planta vacuum infiltration methods that circumvent labor-intensive tissue culture steps entirely, thereby democratizing access to sophisticated legume genetic engineering for diverse research applications.
R108-1 offers superior regeneration capacity and faster tissue culture cycling (ideal for CRISPR and high-throughput work), while A17 serves as the reference genome standard but requires more precise culture conditions. We recommend R108-1 for initial proof-of-concept and A17 for publication-grade reference line studies.
Properly optimized protocols preserve symbiotic competence. Our regeneration media formulations and rooting strategies specifically maintain the developmental plasticity required for rhizobial infection. We routinely verify nodulation capacity in T0 lines before delivering to clients.
Cotyledon-based transformation (bypassing callus) offers the fastest turnaround (direct shoot organogenesis) for A17. Leaf discs from R108-1 provide high efficiency for traditional somatic embryogenesis. Cell suspensions are optimal for transient expression or bioreactor-scale protein production.
Our transient expression service using mesophyll protoplasts or cell suspensions provides functional data (protein localization, promoter activity, CRISPR efficiency) within 2–4 days. This allows rapid iteration before committing to the 6–8 month stable transformation timeline.
Yes. While R108-1 and A17 are standard, we have developed customized protocols for less responsive lines including modified hormone ratios (cytokinin/auxin balances), enhanced virulence Agrobacterium strains, and vacuum infiltration alternatives for lines recalcitrant to tissue culture.
Standard stable transformation delivers T0 plantlets in 5–6 months. Since nodulation assays can be performed on T0 plants with established root systems, you can expect symbiosis phenotyping data within 7–8 months of project initiation, compared to 10–12 months required for seed-advanced generations.

Unlocking the Basics: What is Genetic Transformation and Why It Matters
CRISPR-Cas9: A Comprehensive Guide to Genome Editing in Plants

Agrobacterium tumefaciens-mediated Tobacco Leaf Disk Transformation
Transformation of Immature Wheat Embryos Using Gene Gun Method
Reference