Professional, Customizable Gossypium hirsutum Genetic Engineering Solutions
Lifeasible stands as an industrial leader in plant biotechnology, delivering comprehensive transformation platforms specifically optimized for upland cotton. Gossypium hirsutum represents one of the world's most economically significant crops, providing natural fiber for textiles, high-quality oil for consumption, and protein-rich meal for livestock. Our specialized services bridge the gap between genetic discovery and agricultural application, offering researchers and AgBio professionals a direct pathway from sequence to stable transgenic lines.
Through our established expertise in agricultural genetic engineering, we support diverse research objectives—from enhancing fiber elongation and strength to engineering resistance against bollworm, aphids, and environmental stressors like drought and salinity. Commercial deployment of insect-resistant traits, such as Cry protein expression, demonstrates the transformative potential of cotton biotechnology that we make accessible to our partners.
TARGET GENOTYPES
Coker Series, Acala, etc.
Elite breeding lines and model varieties
TYPICAL YIELD
10–25
Independent T0 Events
EDITING EFFICIENCY
Up to 75%
CRISPR/Cas9 Knockout Efficiency
LEAD TIME
6–9 Months
Vector to T1 Seeds
Standard Package
Efficiency Focused
Premium Package
Full-Service Custody
Stable nuclear integration forms the foundation of heritable cotton improvement. Our optimized Agrobacterium-mediated transformation protocols maximize T-DNA incorporation while minimizing backbone carryover and multicopy insertions that can trigger transgene silencing. For researchers requiring organelle engineering, we additionally offer chloroplast transformation consultation.
![]()
Explant Selection
Preparation of surface-sterilized hypocotyl segments, cotyledonary petioles, or shoot apex tissues; initiation of friable embryogenic callus with optimized auxin ratios.
![]()
Infection & Co-cultivation
Vacuum-assisted Agrobacterium infiltration or standard co-cultivation on acetosyringone-supplemented media to enhance T-DNA transfer.
![]()
Stringent Selection
Progressive hygromycin or kanamycin selection pressure to eliminate chimeric tissue while preserving regenerative capacity of transgenic sectors.
![]()
Regeneration
Controlled somatic embryogenesis transitioning through globular, torpedo, and cotyledonary stages under precise hormone regulation.
![]()
Acclimatization
Gradual hardening from humidified growth chambers to standard greenhouse conditions for robust T0 establishment.
When experimental timelines demand immediate results, our protoplast and rapid-assay systems bypass the 6–9 month regeneration cycle. These platforms enable hypothesis testing, vector validation, and regulatory element characterization within days.
![]()
Vector Design
Optimization of constitutive (CaMV35S, FMV) and tissue-specific (fiber-specific E6, Fb late) promoters for cotton cellular contexts.
![]()
Target Material
High-viability protoplast isolation from embryogenic suspension cultures or mesophyll tissues.
![]()
DNA Delivery
PEG-calcium mediated transfection optimized for large plasmids, or electroporation for high-throughput screening.
![]()
Analysis
Fluorescence microscopy, flow cytometry, quantitative PCR, and protein extraction for rapid expression confirmation.
Lifeasible deploys refined methodologies addressing the unique cell wall composition and regeneration requirements of Gossypium species.
Utilizing disarmed Agrobacterium tumefaciens strains (EHA105, GV3101, AGL1) with acetosyringone induction, we achieve high-efficiency T-DNA transfer into hypocotyl-derived callus. In planta alternatives via wounded shoot apical meristem (SAM) infection eliminate tissue culture requirements for specific applications.
Helium-driven biolistic delivery of gold-particle-coated DNA into embryogenic callus for cultivars exhibiting Agrobacterium resistance. Suitable for high-molecular-weight constructs and organelle targeting.
| Category | Requirements |
| Sample Type | Certified mature seeds, fresh hypocotyl segments (7–10 day seedlings), established embryogenic callus lines, or sterile in vitro plantlets |
| Sample Amount | Minimum 100g mature seeds (~3,000–4,000 seeds) or 30–50g fresh embryogenic callus |
| Pre-Treatment | Seeds debearded, delinted, surface-sterilized; no chemical treatment post-harvest; full cultivar pedigree documentation |
| Storage | Seeds at 4°C, <12% moisture; callus at 25°C on maintenance media; avoid freeze-thaw for viable tissue |
| Shipping | Ambient temperature for seeds with desiccant; cold chain (4–10°C) for living tissue; phytosanitary certificates for international shipments |
| Metadata | Fiber quality parameters (if known), generation/purity, previous transformation history, target locus details, selectable marker preferences |
| Vectors | Full sequence maps, promoter identities, selectable markers, reporter genes; sequence confirmation data |
Enhance core transformation with specialized downstream capabilities:
Molecular Characterization
Copy number determination via southern blot, transcript quantification via RT-qPCR, and western blot protein analysis.
CRISPR/Cas9 Off-Target Screening
Whole-genome NGS scanning for unintended editing events.
Vector Engineering
Multi-gene cassette assembly, synthetic promoter development, and dicot codon optimization.
Subcellular Imaging
Confocal microscopy of fluorescent fusion proteins.
Stress Phenotyping
Controlled environment screening for drought, salt, heat, and pathogen resistance.
Vector Construction
Explant Establishment
Transformation & Selection
Regeneration & Hardening
Molecular Analysis
Seed Production
Note: Extended timelines apply for recalcitrant genotypes or complex multiplex editing projects.

High-Efficiency Agrobacterium-Mediated Transformation Pipeline for Elite Cotton Varieties
Internal project report documenting the successful establishment of a robust Agrobacterium tumefaciens-mediated transformation workflow for upland cotton elite breeding lines. The protocol demonstrates complete regeneration capacity from hypocotyl explants through to fertile T0 plants suitable for field evaluation. The methodology encompasses a six-stage pipeline:
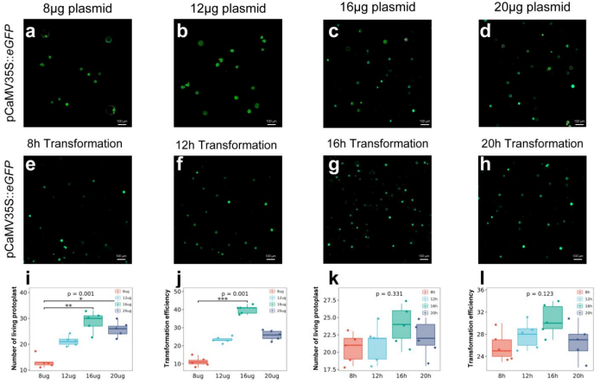
Advanced Protoplast Systems for Rapid Cotton Gene Function Analysis
Research published in International Journal of Molecular Sciences demonstrates highly efficient protocols for cotton callus protoplast isolation and transformation. The methodology achieves superior viability rates through refined enzymatic digestion cocktails tailored to cotton cell wall composition (high pectin and lignin content).
Key technical achievements include optimized osmotic stabilizers maintaining protoplast integrity during isolation from friable callus; PEG-mediated transfection delivering exogenous DNA with high efficiency validated through GFP fluorescence; 24–48 hour turnaround from tissue harvest to expression analysis; and suitability for plasmids up to 20kb including CRISPR/Cas9 cassettes and dual-luciferase reporters.
This system enables immediate validation of gene constructs, promoter characterization, and protein localization studies without the resource investment of stable transformation. The methodology has been applied to fiber development gene validation and stress-response pathway mapping.
Our commitment to precision and reliability has established Lifeasible as a trusted partner for academic and industrial cotton researchers globally:
"Lifeasible successfully transformed our elite Acala breeding line when three other service providers failed. Their team developed a modified biolistic protocol that generated 8 positive T0 lines with single-copy insertions confirmed by Southern blot. The hardening survival rate exceeded 90%, and we now have stable Bt lines entering regional trials."
Dr. K. Richardson
Senior Cotton Geneticist
USA
"We utilized the transient protoplast system to validate 12 CRISPR guide RNAs targeting fiber biosynthesis genes. Within one week, we identified the 4 most efficient guides that showed >60% indel rates. Moving only these candidates to stable transformation saved us approximately 8 months and significant tissue culture costs."
Dr. M. Chen
Principal Investigator, Fiber Genomics
USA
"The Premium Package delivered exactly what we needed for our patent application—T2 homozygous lines with single-copy transgenes, comprehensive molecular characterization, and detailed growth documentation. Their understanding of cotton fiber development biology helped us optimize our construct design before transformation began."
Dr. A. Patel
Director, Trait Development
UK
Cotton-Specific Expertise
Two decades of focused Gossypium transformation experience, encompassing both model lines and commercial elite varieties.
Precision Editing
Advanced CRISPR/Cas9 and Cas12a platforms with on-target efficiency monitoring and off-target screening.
Regulatory Compliance
Operations conducted in certified biosafety facilities meeting international phytosanitary and GMO handling standards.
End-to-End Integration
Seamless transition from transformation through analytical services to breeding support.
Ready to transform your cotton research?
Our technical experts are available to discuss your project requirements, from vector design to greenhouse management. From CRISPR-based gene editing to stable transgenic line development, Lifeasible is your trusted partner for every stage of cotton genetic engineering.
Gossypium hirsutum possesses an allotetraploid genome (AADD, ~2.5 Gb) resulting from ancient hybridization between diploid progenitors. This genomic architecture presents unique challenges for precise editing—homeologous genes (duplicated copies in A and D subgenomes) require careful guide RNA design to avoid off-target effects. Our bioinformatics pipeline specifically addresses subgenome specificity in CRISPR design.
Unlike model plants such as Arabidopsis, cotton relies on somatic embryogenesis for plant regeneration rather than direct organogenesis. This process—converting somatic callus cells into bipolar embryo structures capable of germinating into whole plants—is genotype-dependent and hormone-sensitive. Success requires precise balance of auxin (2,4-D) and cytokinin ratios during the induction phase, followed by gradual hormone withdrawal during maturation. Our proprietary media formulations have optimized these transitions across diverse genetic backgrounds.
Since the initial deployment of Bt cotton in 1996, transgenic varieties have revolutionized agricultural practices in major producing nations. Current stacked-trait varieties combine insect resistance with herbicide tolerance and occasionally drought tolerance modules. The transformation efficiency and molecular characterization rigor provided by service providers directly impacts the speed at which academic discoveries transition to field-ready varieties. Understanding intellectual property landscapes and regulatory data requirements is essential for commercial deployment—a knowledge base integrated into our Premium Package workflows.
Cotton transformation remains highly genotype-dependent. Model varieties like Coker 312 and TM-1 achieve 30–50% callus induction and 15-25% transformation efficiency. Elite Acala and Pima (Egyptian) varieties often exhibit recalcitrance requiring modified hormone regimes or alternative delivery methods. We recommend feasibility assessments for untested varieties.
Our optimized protocols target low-copy events (1–2 insertions) in 60–70% of transgenic lines. Southern blot analysis in our Premium Package confirms copy number, facilitating selection of lines for breeding that show stable Mendelian segregation without silencing.
Yes. We maintain vector collections containing fiber-specific promoters (E6, FbL2A, Exp1) active during elongation and secondary cell wall synthesis phases. These enable temporal-specific expression of transgenes without vegetative tissue metabolic burden.
Editing efficiency varies by target locus but typically ranges 40–75% for single-guide designs. Multiplex editing (2–3 targets) achieves 20–40% simultaneous modification rates. We recommend protoplast pre-screening to identify high-efficiency guides before stable transformation investment.

Pollen Tube Pathway-Mediated Cotton Transformation

SSR Identification of Cotton Salt Tolerance

Transformation of Cucumber

Stable vs. Transient Transformation
Reference