There are four major groups of genetic markers: morphological markers, biochemical markers, cytological markers, and molecular markers. Lifeasible, as a leading plant biotechnology company, has rich experience in the discovery and development of DNA markers, for plant breeding and biodiversity studies.
DNA markers are small regions of DNA sequences that show polymorphism (base deletion, insertion, and substitution) among different individuals. They usually are closely associated with the target genes, and can therefore serve as flags for target gene selection. DNA markers play important roles in the construction of genetic maps, analysis of genetic diversity, map-based cloning, as well as marker-assisted breeding. For the exploitation of genetic markers, several marker systems have been developed, such as restriction fragment length polymorphism (RFLP), random amplification of polymorphic DNA (RAPD), amplified fragment length polymorphism (AFLP), sequence characterized amplified region polymorphism (SCARP), sequence tagged sites (STS), simple sequence repeats (SSR), Inter simple sequence repeat (ISSR), restriction-site associated DNA (RAD), sequence related amplified polymorphism (SRAP), single nucleotide polymorphism (SNP), and so on.
Lifeasible provides marker discovery and development services for a wide variety of plant species, including field crops, vegetables, and ornamentals. We have developed multiple advanced marker discovery methods adapting to the plant system, including:
With our well-established sequencing platforms (FLX Titanium, Solexa, Illumina MiSeq and HiSeq2500, SOLiD, etc.), as well as excellent experts in genetics, statistics, and bioinformatics, Lifeasible offer plant genetic marker discovery services of high accuracy and efficiency, with competitive prices. We are devoted to helping you find the optimal DNA markers for your research.
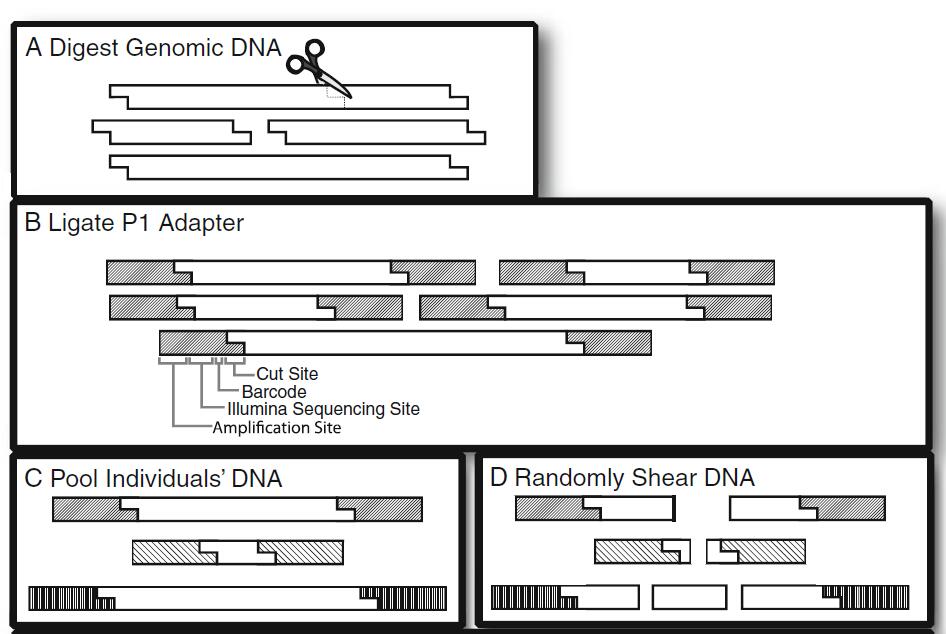
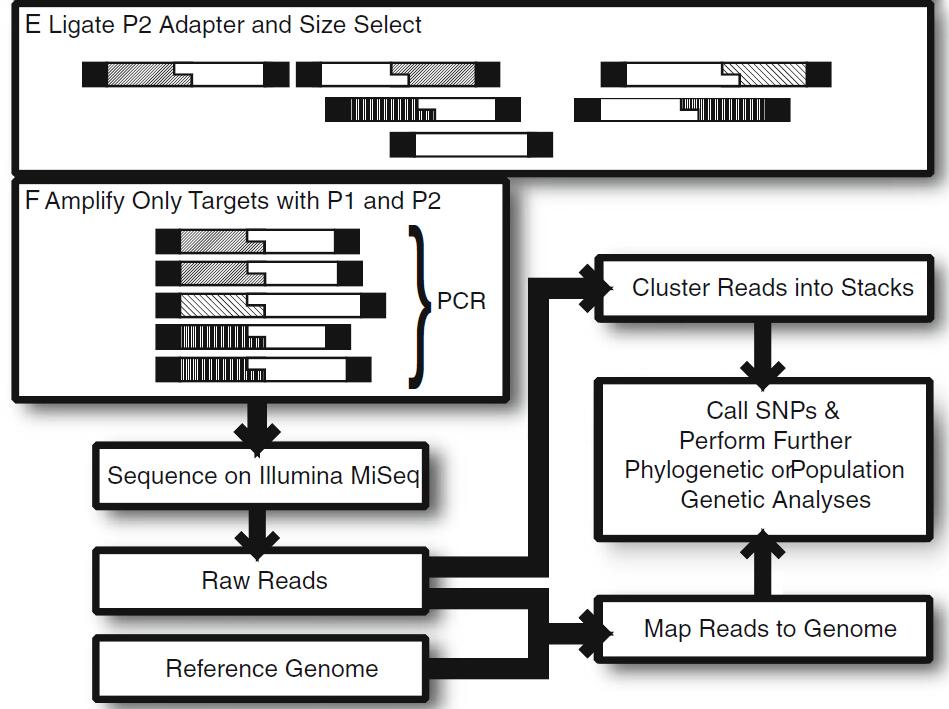 Figure 1. An overview of the workflow for RAD-seq library creation (Bergey et al., 2013)
Figure 1. An overview of the workflow for RAD-seq library creation (Bergey et al., 2013)
Reference: