Wheat is a major staple crop worldwide, but its yield is continuously threatened by diseases such as powdery mildew. Breeding and utilizing disease-resistant genes is the most economical and environmentally friendly strategy to combat these diseases. Plant resistance typically relies on immune receptors encoded by resistance (R) genes. These receptors recognize avirulence (Avr) effector proteins secreted by pathogens, thereby activating an immune response, usually manifested as programmed cell death. In wheat, most known R genes encode NLR-type receptors, but in recent years, a new type of immune receptor called kinase fusion protein (KFP) has been discovered, and its mechanism of action is still unclear. The Pm4 gene is a typical KFP, which not only confers broad-spectrum resistance to powdery mildew in wheat but also resists devastating rice blast disease, making it highly valuable in breeding. However, pathogens can evolve corresponding virulence to overcome Pm4 resistance, and the molecular mechanism of this process is a cutting-edge issue in plant pathology research. Understanding this process is crucial for understanding KFP-mediated immune mechanisms and achieving durable disease resistance breeding.
On January 12, 2026, a team from the University of Zurich, led by Javier Sánchez-Martín and Beat Keller, published a research paper in Nature Plants titled "Virulence on Pm4 kinase-based resistance is determined by two divergent wheat powdery mildew effectors." This study revealed the complex molecular mechanism by which wheat powdery mildew overcomes Pm4 resistance, not through traditional avirulence effector mutations, but through a sophisticated dual-effector system. The study first identified and validated the avirulence effector AvrPm4, which is directly recognized by the Pm4 receptor. Subsequently, through population genetic analysis, they discovered an independent suppressor effector, SvrPm4, which effectively suppresses the immune response triggered by the AvrPm4-Pm4 interaction. Even more surprisingly, this suppressor SvrPm4 is itself an avirulence effector recognized by another NLR-type resistance gene, Pm1a. This finding reveals complex synergistic interactions between different types of immune receptors, providing a completely new perspective for designing durable disease resistance breeding strategies.
To identify the effector recognized by Pm4, the research team used UV mutagenesis (AvrXpose) on a powdery mildew strain that could not infect Pm4 wheat, and screened for six acquired virulence mutants that could successfully infect Pm4a and Pm4b wheat. Through whole-genome sequencing of these mutants, they found that all six mutants had loss-of-function mutations in the same gene (Bgt-55142). This indicated that Bgt-55142 is the avirulence gene recognized by Pm4, and the researchers named it AvrPm4.
Structural analysis showed that AvrPm4 is an atypical powdery mildew effector. It is much larger than other known Avr proteins, and its protein structure exhibits chimeric characteristics: the N-terminus is a typical RNase-like domain, while the C-terminus is an MEA domain rich in repetitive sequences with an unknown function. Interestingly, this MEA domain has a highly similar sequence motif to the rice blast fungus effector AVR-Rmg8, which is also recognized by Pm4, suggesting that this domain plays a key role in the recognition by Pm4. To verify the function of AvrPm4, the researchers transiently expressed AvrPm4 in wheat protoplasts containing the Pm4b gene, successfully inducing cell death. Expression of truncated or mutated versions of avrPm4 from the virulence mutants did not induce cell death, functionally confirming that AvrPm4 is the target recognized by Pm4.
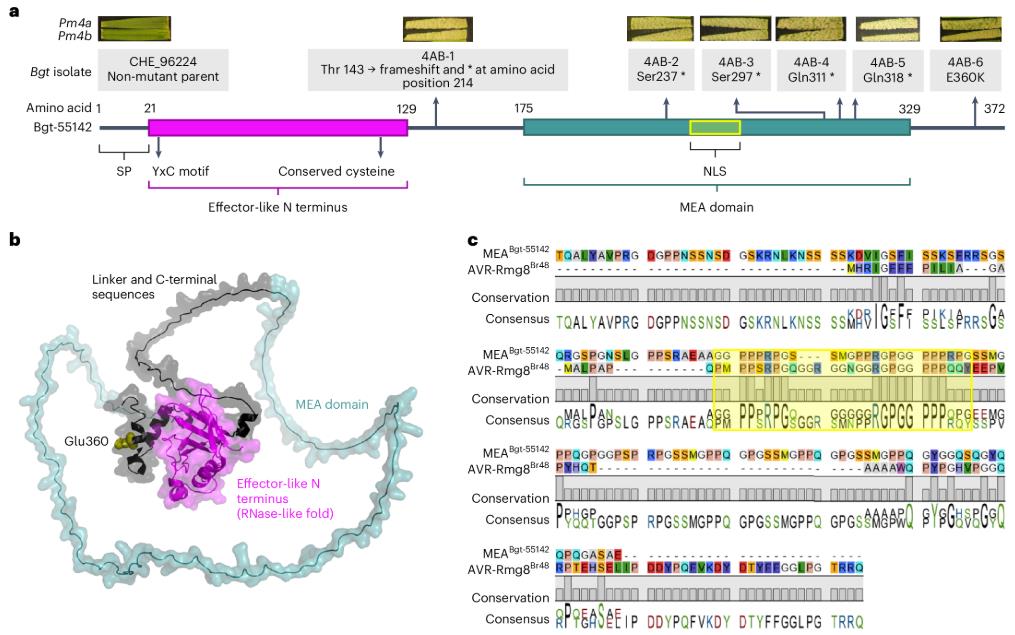
Figure 1. Pm4 gain-of-virulence mutants all exhibit mutations in Bgt-55142, encoding a non-canonical effector. (Bernasconi, et al. 2026)
The Pm4 gene encodes a kinase fusion protein, theoretically possessing kinase activity. To investigate its recognition mechanism, the research team demonstrated through split-luciferase complementation experiments (split-LUC) that the AvrPm4 protein directly interacts physically with both splice variants of Pm4b (Pm4b-V1 and Pm4b-V2) within plant cells. Subcellular localization experiments further showed that AvrPm4 co-localizes with Pm4b-V1 in the cytoplasm, and when co-expressed with Pm4b-V2, which is localized to the endoplasmic reticulum, AvrPm4 is recruited to the endoplasmic reticulum, further confirming their close interaction.
In vitro kinase activity experiments showed that the Pm4 protein itself has autophosphorylation activity and can directly phosphorylate the AvrPm4 protein. The study also found that the kinase activity of Pm4 is crucial for its disease resistance function; Pm4 mutants with inactivated kinase activity cannot mediate the immune response. Interestingly, even a truncated version of avrPm4 that does not induce cell death can be phosphorylated by Pm4. This result indicates that phosphorylation is a necessary step for Pm4 to recognize AvrPm4 and activate immunity, but phosphorylation alone is not sufficient to trigger a complete immune response.
In nature, many powdery mildew strains capable of infecting Pm4 wheat have intact AvrPm4 gene sequences, unlike the virulence mutants induced in the laboratory. This phenomenon suggests a more complex virulence mechanism. To uncover this mechanism, the research team conducted genome-wide association studies (GWAS) and QTL mapping on 78 powdery mildew strains worldwide. Surprisingly, they found that the key genetic locus determining powdery mildew virulence against Pm4 was not on chromosome 5, where AvrPm4 is located, but in a completely new region on chromosome 8.
The researchers hypothesized that this region might encode a suppressor effector that interferes with Pm4's recognition of AvrPm4 or downstream signal transduction. By screening candidate effector genes in this region, they identified a gene called Bgt-51526. Functional experiments confirmed that when Bgt-51526 from a virulent strain (named SvrPm4) was co-expressed with AvrPm4 and Pm4, the cell death induced by the AvrPm4-Pm4 interaction was significantly suppressed, while the Bgt-51526 allele from a non-virulent strain (named svrPm4) did not have this suppressive function. This clearly demonstrates that powdery mildew virulence against Pm4 is achieved through a two-component system: the pathogen retains AvrPm4 while evolving a functional suppressor, SvrPm4, to circumvent Pm4's immune surveillance.
The research team discovered that this suppressor, SvrPm4 (Bgt-51526), is not a new finding; it had previously been identified as AvrPm1a, an avirulence effector recognized by another wheat disease resistance gene, Pm1a (a typical NLR receptor). This study verified this finding, showing that in Nicotiana benthamiana, the functional suppressor SvrPm4 is recognized by Pm1a and triggers a strong hypersensitive response (cell death), while the non-suppressive svrPm4 is not recognized by Pm1a.
This discovery reveals the dual identity and function of the SvrPm4 effector: when the host wheat carries the Pm4 gene, SvrPm4 acts as a virulence factor, helping the pathogen evade immunity; however, when the host carries the Pm1a gene, SvrPm4 becomes an avirulence factor, leading to the pathogen's recognition and elimination. This function switching, dependent on the host genotype, vividly demonstrates the complexity and dynamism of co-evolution between pathogens and hosts.
This study systematically reveals the ingenious molecular strategy used by wheat powdery mildew to overcome the KFP-type resistance gene Pm4, namely the "recognition-suppression" dual effector model. This finding not only deepens our understanding of kinase fusion protein-mediated plant immune signal transduction but also provides new insights into the evolution of pathogen virulence.
The most important contribution of this research is the revelation of the dual function of the SvrPm4 effector: it is both a suppressor of Pm4 and a recognition target of Pm1a. This unique biological phenomenon provides a highly attractive evolutionary trap strategy for crop disease resistance breeding. By combining Pm4 (KFP-type) and Pm1a (NLR-type), two different types of resistance genes, in the same wheat variety, a dilemma can be created for powdery mildew: to overcome Pm4 resistance, the pathogen must maintain functional SvrPm4; however, once it possesses SvrPm4, it will be recognized by Pm1a and unable to survive. This breeding strategy, which forces the pathogen to make a fatal choice between two resistance pressures, is expected to build more durable and stable field resistance, providing strong theoretical support and technical pathways for ensuring safe wheat production.