Some Cas proteins, such as Cas12a (also known as Cpf1), cleave surrounding non-specific single-stranded DNA after cleaving the target DNA strand. This phenomenon is called trans-cleavage. CRISPR-Cas12a has been widely used in molecular diagnostics due to its highly efficient trans-cleavage activity. For the well-known Cas9 protein, its most common use is the specific cis-cleavage of target double-stranded DNA guided by guide RNA. As early as May 2024, Nature Biotechnology published a research paper titled "Trans-nuclease activity of Cas9 activated by DNA or RNA target binding," which found that target DNA activated the trans-cleavage activity of Cas9-sgRNA, and that using a tracrRNA (trans-activating crRNA)-crRNA dual-guide system could significantly enhance the trans-cleavage activity of Cas9, leading to non-specific cleavage of poly T/C/A molecules and ssDNA, indicating its potential application in molecular diagnostics.
On January 24, 2026, a research team from CSIC – University of Valencia, led by Guillermo Rodrigo, published a research paper in Nature Communications titled "CRISPR-Cas9 trans-cleavage is hindered by a flanked R-loop, an elongated spacer, and an inactive HNH domain." The researchers found that Cas9's trans-cleavage is not a random phenomenon. When an unflanked R-loop is formed on the RNA 5' end side, when the spacer length in sgRNA exceeds the classic 20 bp, and when an inactive HNH domain is used, the trans-cleavage activity of Cas9 is significantly weakened, thus allowing for precise control of Cas9's cleavage activity.
The researchers used dsDNA of different lengths as targets to investigate the effect of the R-loop (RNA-DNA hybrid) RNA 5' end flanking structure on trans-cleavage activity. When the target was short dsDNA (short pre-PAM region, such as 20 bp), an unflanked R-loop was formed, and Cas9 exhibited high trans-cleavage activity; however, when the target was long dsDNA (long pre-PAM region, such as 36 bp), the R-loop was flanked by double-stranded DNA at the 5' end, and the trans-cleavage activity was significantly reduced. This indicates that the R-loop structure at the RNA 5' end is crucial for Cas9 trans-cleavage activity.
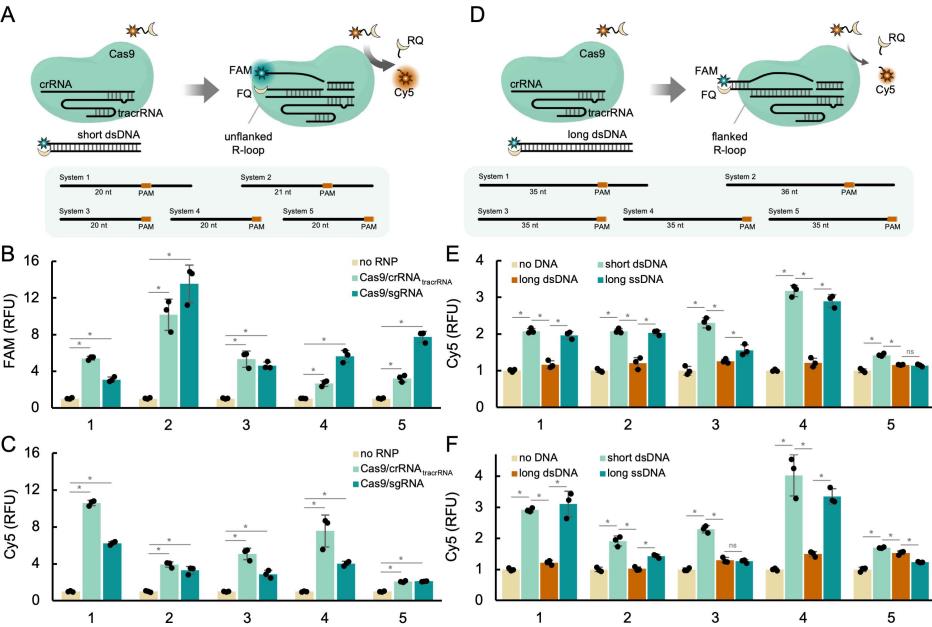
Figure 1. The R-loop structure is crucial for the trans-cleavage activity of Cas9. (Montagud-Martínez, et al. 2026)
The researchers tested the effect of different gRNA spacer lengths (19-24 nt) on Cas9 trans-cleavage activity. They found that when the spacer length exceeded the classic 20 nt, such as 22 and 24 nt, the trans-cleavage activity decreased significantly, by 49% and 79%, respectively, and the trend of decreasing trans-cleavage activity with increasing spacer length showed a linear correlation. Using a 24 nt spacer did not significantly affect Cas9's target binding and cis-cleavage ability, indicating that spacer length mainly affects Cas9's trans-cleavage activity. The elongation of the spacer may alter the interaction between the gRNA 5' end and Cas9, thus affecting its conformational stability. Furthermore, the trans-cleavage activity of Cas9 exhibits higher sensitivity and sequence dependence to base mismatches in the spacer region.
In a practical application context, the researchers designed amplicons with different lengths of the pre-PAM region, targeting SARS-CoV-2 RNA. The results showed that only amplicons with a short pre-PAM region (forming an unflanked R-loop) could be detected, while amplicons with a long pre-PAM region could not, further confirming the importance of the R-loop flanking structure for CRISPR-Cas9 diagnostic applications. Analysis of the complex structure of Cas9 with different lengths of target DNA revealed that when the R-loop is flanked, the solvent accessibility of the RuvC active site is reduced, potentially limiting its cleavage of the ssDNA probe.
This study systematically revealed three key structural factors that regulate Cas9 trans-cleavage activity: R-loop flanking structure, spacer length, and HNH domain activity. The research findings not only deepen our understanding of the mechanisms underlying the multifaceted functions of Cas9, but also provide important design and guidance principles for optimizing the application of CRISPR-Cas9 in nucleic acid detection and diagnosis.