The growth and development of all organisms depend on the coordinated regulation of gene expression in time and space, which is largely controlled by non-coding sequences in the genome.
The main challenge of genomics to promote crop improvement is the functional annotation of cis-regulatory elements in crop genomes and the ability to use these sequences to fine-tune target pathways through breeding or biotechnology.
Recently, researchers such as Dr. Andrea Eveland discovered that in the early developmental window of maize, non-coding “functional” genome maps are essential for pollen formation.
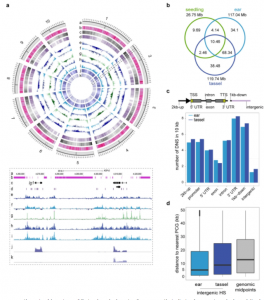
In this study, the researchers integrated information about chromatin structure, transcript profiles, and genome-wide association studies. Their analysis provided a comprehensive understanding of the regulation of inflorescence differentiation in major cereal crops.
Related results were published in the “Genome biology”.
Over the past century, hybrid-based corn breeding and improvement has led people to choose smaller ears, which will intercept less light, use fewer resources, and produce higher yields.
However, since ears and maize are developed through a common developmental program, further improvement of maize traits will require decoupling the program. Understanding how the same genes are regulated differently in ears and maize, and using this specificity to control a difference between each other will enhance maize breeding.
Eveland's research focuses on the developmental mechanisms that control the structural characteristics of cereal crops. Specifically, she studied how stem cells form plant organs, and how mutations in the underlying gene regulatory network precisely regulate plant morphology. Her team integrated computational and experimental methods to explore how the perturbation of these gene networks can change the morphology of the species and finally determine the target of modification.
Reference
Parvathaneni, R. K., et al. (2020) The regulatory landscape of early maize inflorescence development. Genome Biology. doi.org/10.1186/s13059-020-02070-8.