Lifeasible is a specialized biotechnology company in plant transgenesis. Transcription activator-like effector nucleases (TALENs), highly efficient and versatile tools for genome editing, have been extensively used in plant gene function research as well as plant breeding. The technology exhibits reduced off-target effects and cytotoxicities and simple protein–DNA code compared with zinc finger nucleases (ZFNs).
TALENs consist of TALE DNA-binding domains and cleavage domains of restriction endonuclease FokI. The DNA binding domains are composed of a number of tandem repeats varying from 1.5-33.5 typically. Each repeat contains 34 amino acids with divergent 12th and 13th amino acids (termed as “repeat variable diresidues” (RVDs)) that specifically recognize one nucleotide. Briefly, the decoding mechanism of RVDs is that NI (Asn Ile), HD (His Asp), NN (Asn Asn) and NG (Asn Gly) recognizes A, C, G, T, respectively.
The DNA cleavage domain causes DNA double strand breaks (DSBs) in dimers by a sequence-independent way. The DSBs are repaired by either non-homologous end joining (NHEJ) or homology directed repair (HDR), resulting in several types of genomic alterations in the broken sites such as point mutations, deletions, insertions, inversions, duplications, and translocations.
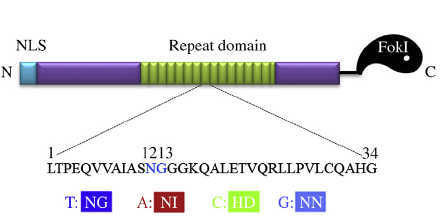 Figure 1. Structure of transcription activator-like effector nucleases (Wei et al., 2013)
Figure 1. Structure of transcription activator-like effector nucleases (Wei et al., 2013)
Lifeasible has extensive experience in customized TALEs such as restriction enzyme and ligation (REAL), gold gate PCR (GG-PCR), GG-Vector, fast ligation-based automatable solid-phase high-throughput (FLASH), iterative capped assembly (ICA), ligation-independent cloning (LIC) and unit assembly (UA). We also can make modified FokI domains for heterodimers to improve the specificity of cleavage. At Lifeasible, a large selection of model plant transformation systems such as Arabidopsis, rice, wheat, alfalfa, corn, tobacco, cotton and others are readily available to accommodate the diverse needs of the clients.
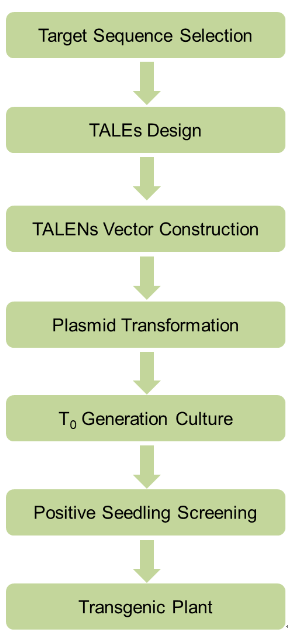
Experimental process:
Reference:
Lifeasible has established a one-stop service platform for plants. In addition to obtaining customized solutions for plant genetic engineering, customers can also conduct follow-up analysis and research on plants through our analysis platform. The analytical services we provide include but are not limited to the following:
STU-CRISPR System Improves Plant Genome Editing Efficiency
April 19, 2024
Application of Exosomes in Facial Beauty
April 12, 2024