Baculoviruses are a family of large, double-stranded DNA insect viruses with a genome size of approximately 134 kb. They not only efficiently express recombinant proteins within insect cells but can also transduce various mammalian cell types—including HEK293T cells, induced pluripotent stem cells (iPSCs), and hepatocellular carcinoma cells—to achieve transient gene expression. Leveraging these characteristics, baculoviruses have emerged as critical gene delivery vectors in the fields of cancer gene therapy, regenerative medicine, genome editing, and prime editing.
However, the generation of traditional recombinant baculoviruses relies on inefficient homologous recombination. While existing improved systems—such as flashBAC and BaculoDirect—have enhanced recombination efficiency, they depend on proprietary linear genomes, making large-scale or multiplex viral construction prohibitively expensive. Although the Bac-to-Bac system offers a cost-effective alternative, it is restricted to a single attTn7 integration site, and exogenous genes are prone to loss during successive viral passages. These technical bottlenecks have limited the application of baculoviruses in complex genetic engineering endeavors.
To overcome these existing technical limitations, a research team has developed the SHOT 2.0 system. This system is based on Type V-K CRISPR-associated transposon technology, utilizing RNA guidance to achieve site-specific integration without inducing DNA double-strand breaks.
The core of the system comprises two classes of plasmids:
In the initial version, SHOT 1.0, approximately 28.6% of the baculovirus genomes could be edited; however, a substantial proportion of unedited genomes remained. Through systematic optimization, the research team discovered that the primary factor limiting editing efficiency was not the copy number of the donor plasmid, but rather the expression level of the sgRNA. Consequently, SHOT 2.0 incorporates an additional sgRNA expression cassette into the donor plasmid and utilizes a high-copy-number replicon; simultaneously, the replicon in the helper plasmid is replaced with a compatible p15A replicon.
The editing efficiency of the optimized system has been boosted to 79.4%. When combined with a colony streaking purification step, residual unedited bacmids at the target locus were undetectable after just two rounds of streaking, yielding strains with 100% editing purity. SHOT 2.0 successfully facilitates the efficient integration of large DNA fragments—at least 14 kb in size—far exceeding the practical limit of 3–4 kb associated with traditional recombinant engineering techniques.
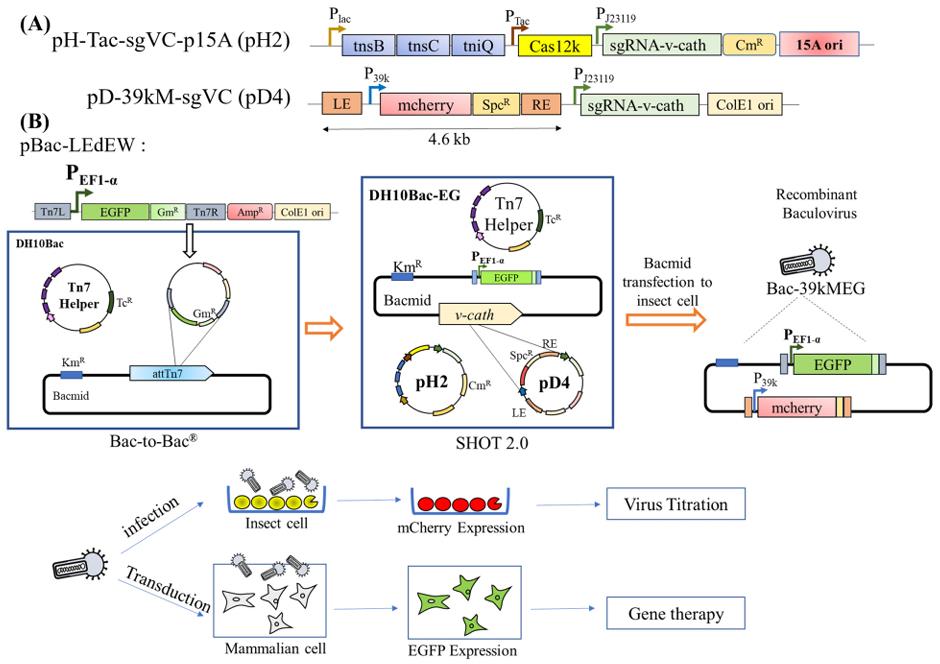
Figure 1. SHOT 2.0 is compatible with Bac-to-Bac for facile baculovirus engineering. (Dang, et al. 2026)
Another key advantage of SHOT 2.0 is its complete compatibility with existing Bac-to-Bac workflows. The research team demonstrated an orthogonal dual-gene insertion strategy:
The resulting recombinant baculovirus replicated efficiently within insect cells, achieving a viral titer of 108–109 pfu/mL—a level comparable to that of the standard Bac-to-Bac system. Following transduction into HEK293-FT cells, 97.2% of the cells expressed EGFP, while RFP expression remained negligible; this result confirmed the insect-cell-specific activity of the 39k promoter, which remains largely inactive in mammalian cells. This dual-locus strategy offers a novel approach for simultaneously delivering both reporter genes and therapeutic genes.
During serial passaging, exogenous genes integrated into the attTn7 locus of baculoviruses are prone to progressive loss—a critical issue that currently constrains the large-scale application of these systems. Previous studies have demonstrated that relocating exogenous genes to the ODVe56 locus can significantly enhance genomic stability.
Leveraging the programmable targeting capabilities of SHOT 2.0, the research team was able to redirect the exogenous gene from the v-cath locus to the ODVe56 locus simply by altering the sgRNA sequence. Quantitative analysis following 10 rounds of serial passaging revealed the following:
In terms of transduction efficiency, the 10th-generation ODVe56-targeted virus was still capable of transducing approximately 95% of HEK293T cells, and its mean fluorescence intensity was significantly higher than that of the v-cath-targeted virus. This result confirms that genetic stability in recombinant baculoviruses can be substantially improved through locus selection—a finding of significant importance for applications requiring long-term passaging, such as vaccine production and stem cell engineering.
Prime editing technology enables precise base substitutions, insertions, and deletions without introducing DNA double-strand breaks; however, the efficient delivery of the complete editing system—which exceeds 7 kb in size—has long remained a technical challenge. Common vectors, such as adeno-associated viruses (AAV), are unable to accommodate full-length prime editing components due to packaging capacity limitations, necessitating the adoption of dual-vector strategies that result in reduced efficiency and increased system complexity.
Leveraging the large-fragment integration capabilities of SHOT 2.0, the research team successfully packaged the PE5max prime editing system into a baculovirus. The specific strategy employed was as follows:
The resulting all-in-one baculovirus, designated Bac-PE5max, demonstrated exceptional editing efficiency in HEK293T cells:
In contrast, the efficiency of traditional dual-plasmid transfection systems was markedly lower. Off-target analysis revealed that at three predicted off-target sites exhibiting high sequence homology, the frequency of off-target editing remained below 0.4%, thereby demonstrating that the system maintains excellent specificity while delivering high editing efficiency.
Due to their inherent resistance to transfection, induced pluripotent stem cells (iPSCs) and various liver cancer cell lines have long posed a formidable challenge for gene editing. The all-in-one baculovirus (Bac-PE5max) demonstrated exceptional performance in these hard-to-transfect cells:
In induced pluripotent stem cells, the prime editing efficiency of Bac-PE5max significantly outperformed that of dual-plasmid transfection:
Across four hepatocellular carcinoma cell lines, the efficiency of +1 CTT insertion ranged from 5.2% to 37.1%, with the highest efficiency observed in Hep3B cells. This achievement is comparable to—or even surpasses—that of existing non-viral delivery systems, and far exceeds the performance of dual-vector adeno-associated virus (AAV) strategies in similar cell types.
Baculoviruses inherently possess high transduction efficiency in primary and hard-to-transfect cells; when combined with the programmable genomic integration capabilities of SHOT 2.0, they demonstrate unique advantages for the clinical translation of prime editing applications.
The establishment of SHOT 2.0 provides a flexible and programmable new toolkit for baculovirus engineering. Compared to Cre-recombinase systems that rely on pre-installed loxP sites, the RNA-guided integration method allows users to freely define insertion sites without the need for tedious preliminary modifications to the host genome.
A current limitation of the system is that the pegRNA/sgRNA module remains situated at the relatively unstable attTn7 site, which restricts the number of viral passages. Future improvements could involve integrating orthogonal Type I-F CRISPR-transposon systems or combining the Cre/loxP strategy utilized in the MultiBac system to enable iterative, stable integration at multiple genomic loci. Furthermore, introducing synthetic regulatory circuits to enable time-limited expression of the Cas9 nickase could further mitigate potential off-target risks.
Overall, SHOT 2.0 not only expands the scope of baculovirus applications in basic research but also provides a robust delivery platform for precise genomic editing in the fields of gene therapy and regenerative medicine.