Gene editing technologies are revolutionizing agriculture and medicine; however, traditional tools such as Cas9 are relatively large in size, making their efficient delivery via viral vectors challenging. TnpB is a class of RNA-guided endonucleases—considered the evolutionary ancestor of CRISPR-Cas12—and is regarded as an ideal next-generation editing tool due to its low molecular weight and compact structure. Nevertheless, the editing activity of naturally occurring TnpB is generally weak, which has limited its practical application.
Recently, the teams led by Jennifer A. Doudna and David F. Savage published a study in Nature Biotechnology. By employing deep mutational scanning, they systematically elucidated the sequence-function relationships of ISDra2 TnpB, successfully developing highly active engineered variants that provide a new paradigm for the design of compact genome editing tools.
The research team utilized Deep Mutational Scanning (DMS) technology to establish a positive selection assay for TnpB-mediated DNA cleavage within a yeast system. This method quantitatively assesses the impact of each specific mutation on nuclease activity by monitoring the growth of yeast cells in an adenine-deficient culture medium.
The experimental design involved two independent plasmid libraries:
Each variant was associated with approximately 30 unique barcodes. By calculating the relative enrichment—derived from the ratio of barcode abundance under selective versus non-selective conditions—the researchers achieved high-throughput functional assessment of the variants.
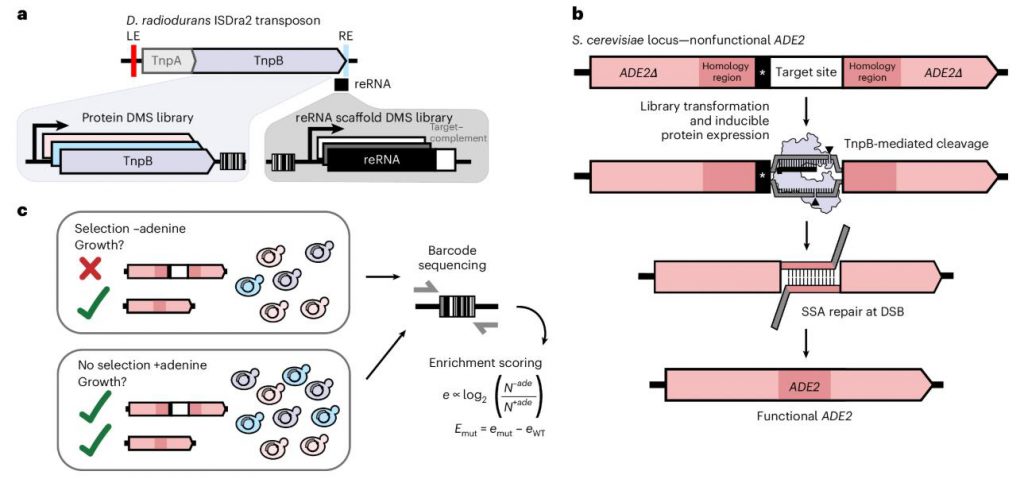
Figure 1. Design of DMS libraries and optimized in vivo selection for endonuclease activity in yeast. (Thornton, et al. 2026)
The reRNA component accounts for approximately half of the molecular weight of the TnpB ribonucleoprotein complex, and its 116-nucleotide scaffold is critical for cleavage activity. The study revealed the following findings:
In EGFP knockout experiments conducted in HEK293T cells, hinge-region deletion variants demonstrated higher editing efficiency than the wild-type; conversely, while Trim2 variants were highly enriched in yeast, their efficiency declined in mammalian cells. These results suggest that Stem 2 plays a pivotal role in regulating the release of the RuvC domain, and that variant activity is differentially influenced by distinct host environments.
The protein DMS data reveals a rich functional landscape:
Of particular note are the following observations:
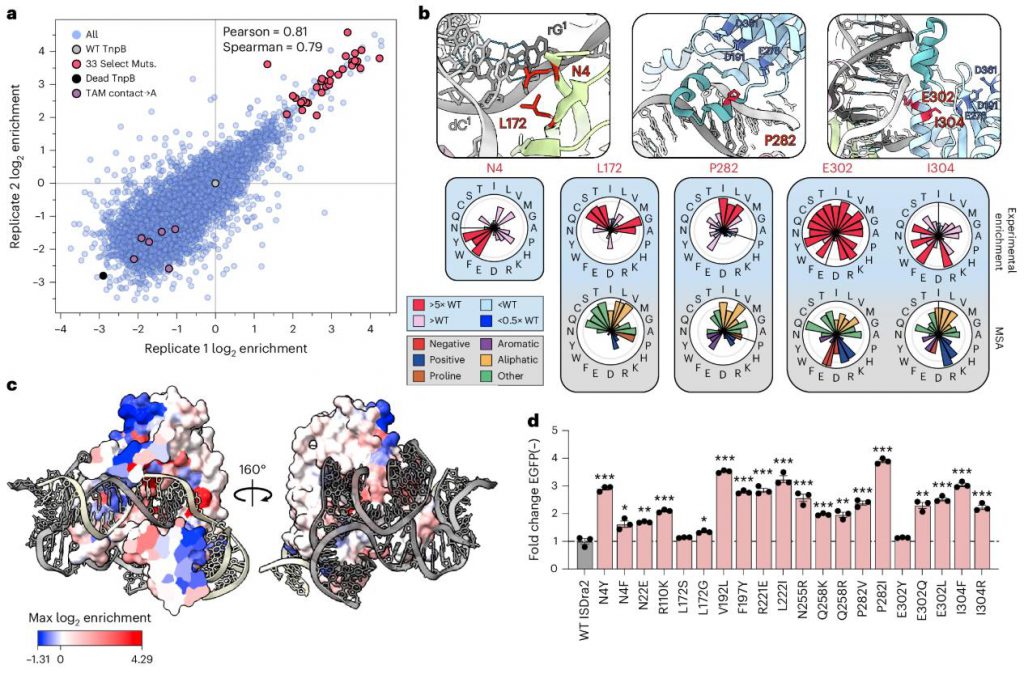
Figure 2. Activating mutations inform mechanistic insights and engineering. (Thornton, et al. 2026)
Based on the DMS data, the research team selected 33 highly enriched single-point mutations spanning 19 distinct sites and constructed a library of approximately 5,000 combinatorial variants using nicking mutagenesis. Following two rounds of yeast-based selection, five highly active combinatorial variants (designated eTnpBa through eTnpBe) were successfully identified. Testing in HEK293T cells revealed the following:
The application of compact editing tools in plants has historically been constrained by delivery efficiency and cargo capacity. This study evaluated the editing capabilities of eTnpBc and eTnpBe (designated as TnpB-KYLI and TnpB-VGIRL, respectively) across various plant species:
Off-target analysis indicated that the engineered variants maintained off-target activity levels comparable to—or even lower than—those of the wild-type in N. benthamiana.
This study elucidates the core elements governing the regulation of TnpB activity:
The study further revealed that activating mutations identified in ISDra2 can be successfully transferred to its homologs—ISYmu1 and ISAba30—despite their low sequence similarity, thereby demonstrating the universality of this activation mechanism across the TnpB family.
The successful engineering of TnpB-KYLI and TnpB-VGIRL marks a pivotal moment, ushering compact gene-editing tools into a phase of high efficiency and practical utility. Its significance is underscored by the following aspects:
The research team had previously demonstrated that TnpB-KYLI, delivered via viral vectors, achieved heritable editing rates exceeding 50% in the progeny of N. benthamiana—a performance significantly superior to that of the wild-type enzyme. With further optimization of delivery methods and the expansion of the TAM recognition repertoire, engineered TnpB variants are poised to emerge as a pivotal platform technology within the field of gene editing.
| Cat# | Product Name | Size |
| ACC-100 | GV3101 Chemically Competent Cell | 100μL/tube |
| ACC-103 | EHA105 Chemically Competent Cell | 100μL/tube |
| ACC-105 | AGL1 Chemically Competent Cell | 100μL/tube |
| ACC-107 | LBA4404 Chemically Competent Cell | 100μL/tube |
| ACC-108 | EHA101 Chemically Competent Cell | 100μL/tube |
| ACC-117 | Ar.Qual Chemically Competent Cell | 100μL/tube |
| ACC-118 | MSU440 Chemically Competent Cell | 100μL/tube |
| ACC-119 | C58C1 Chemically Competent Cell | 100μL/tube |
| ACC-121 | K599 Chemically Competent Cell | 100μL/tube |
| ACC-122 | Ar.A4 Electroporation Competent Cell | 50μL/tube |