Antioxidant Enzymes Activity Analysis
Reactive oxygen species (ROS), such as superoxide radicals (O2−), singlet oxygen (1O2), hydrogen peroxide (H2O2) and hydroxyl radicals (OH−), can accumulate in plant cells under oxidative stresses. ROS can cause damage to lipids, proteins, and DNAs. To cope with this damage, the plant has evolved to be able to produce multiple antioxidant enzymes, which can be indicators for oxidative stresses (Figure 1).
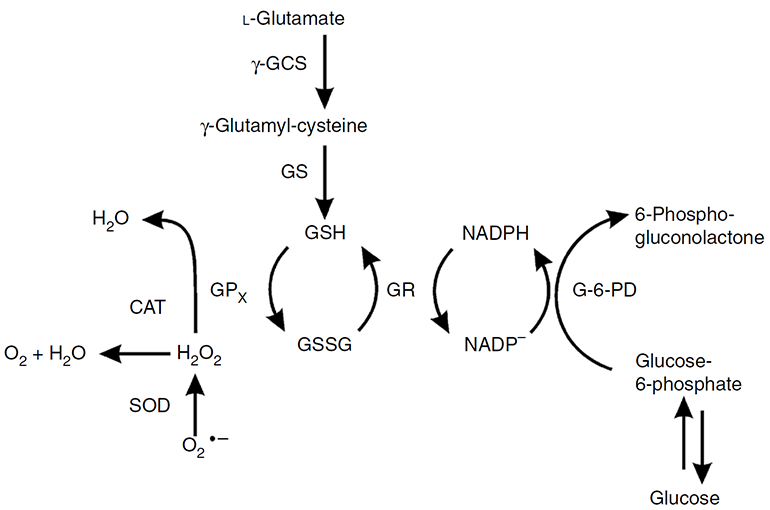 Figure 1. Functions of three antioxidant enzymes--catalase, superoxide dismutase, and peroxidase (represented by glutathione peroxidase (GPx)) (Weydert and Cullen, 2009).
Figure 1. Functions of three antioxidant enzymes--catalase, superoxide dismutase, and peroxidase (represented by glutathione peroxidase (GPx)) (Weydert and Cullen, 2009).
At Lifeasible, we offer a variety of activity analysis for antioxidant enzymes, which include, but not limited to:
Catalase (CAT) activity analysis
CAT catalyzes the decomposition of H2O2 to water and oxygen (O2) (Figure 1). We provide a variety of methods to quantify CAT activity.
- Measurement of the reduction in H2O2 absorbance at 240 nm. This is the most commonly used method for the evaluation of CAT activity. One unit (U) of CAT activity is defined as the amount of enzyme that causes an absorbance change of 0.001 per minute under assay conditions.
- Quantification of the remaining H2O2. Multiple protocols are applicable for this objective, including:
(1) Labeling of the unconverted H2O2 with the fluorogenic probe. The product is measured colorimetrically at 570 nm or fluorometrically at Ex/Em = 535/587 nm.
(2) An enzyme-linked colourimetric detection method which employs 3, 5-dichloro-2-hydroxy-benzenesulfonic acid (DHBS), 4-aminoantipyrine (AAP) and peroxidase. The resulting quinoneimine dye is measured at 520 nm.
(3) Reacting with cobalt (II) in the presence of bicarbonate ions. The concentration of end product carbonato-cobaltate (III) complex ([Co (CO3)3] Co) is assessed at 440 nm.
- Measurement of the formaldehyde produced by CAT with methanol in the presence of H2O2. The formaldehyde is determined with purpald (4-Amino-3-hydrazino-5-mercapto-1, 2, 4-triazole, as a chromogen) and potassium periodate at 550 nm.
- Gas pressure method. The concentration of oxygen gas formed following H2O2 degradation can be measured using an O2 gas sensor.
Superoxide dismutase (SOD) activity analysis
SOD catalyzes the dismutation of superoxide radicals (O2−) to molecular O2 and H2O2 (Figure 2). There are three types of SOD: Cu/Zn SOD (SOD1) in the cytosol, Mn SOD (SOD2) in mitochondria, and EC SOD (SOD3) in extracellular space. We provide various methods for SOD activity analysis.
- NBT, WST-1, and cytochrome c methods. These methods rely on the inhibition of photoreduction of nitroblue tetrazolium (NBT), WST-1, or cytochrome c by SOD. The xanthine-xanthine oxidase system is employed to generate O2−, which can catalyze NBT, WST-1, or cytochrome c and produce a colored product. The concentration of the corresponding product is read at 550, 450, and 550 nm, respectively. One unit of SOD activity is defined as the amount of enzyme causing 50% inhibition of photochemical reduction of NBT, WST-1, or cytochrome c.
- BXT-01050 method. This method is based on the SOD-mediated increase in the autooxidation rate of 5, 6, 6a, 11b-tetrahydro-3, 9, 10-trihydrobenzo[c]fluorene (BXT-01050) in aqueous alkaline solution. The BXT-01050 auto-oxidation yields a chromophore, which has a maximal absorbance at 525 nm. SOD activity is determined by the ratio of the auto-oxidation rates measured in the presence or absence of SOD.
- Oxygen Radical Absorbance Capacity (ORAC) spectrofluorimetric method. This method relies on the inhibition of SOD in the fluorescence production of protein β-phycoerythrin (β-PE), in the presence of free radicals or oxidizing species, which is generated by 2, 2-azobis (2-amidinopropane) dihydrochloride (ABAP). The fluorescence is measured at Ex/Em = 540/565 nm.
- The Spectrophotometric Ferric Reducing Antioxidant Power (FRAP) method. The principle of this method is that in the presence of a suitable oxidizing solution (e.g., FeCl3), SOD can be reduced by obtaining of cation radical from the N, N-dimethyl-pphenylendiamine dihydrochloride (DMPD). The cation radical displays an absorption peak at 514 nm.
Peroxidase (POD) activity analysis
POD is a hemoprotein that catalyzes the oxidation of a number of substrates by H2O2: H2O2 +substrate−H2→substrate+2H2O. At Lifeasible, we provide four solutions for POD activity analysis, based on the specific substrates.
- Pyrogallol method. For this method, Pyrogallol is used as the substrate, and the absorption increase caused by oxidation of pyrogallol by H2O2 is measured at 420 nm spectrophotometrically. One unit of the POD will form 1.0 mg of purpurogallin from pyrogallol in 20 seconds at pH 6.0 at 20 °C.
- 4-methylcatechol method. The POD activity is determined using 4-methylcatechol as a substrate, and the product amount is determined at 420 nm spectrophotometrically. One unit of enzyme activity is defined as 0.001 changes in absorbance per minute under assay conditions.
- 4-aminoantipyrine method. This method employs 4-aminoantipyrine as the hydrogen donor. The reaction rate is determined by measuring the absorbance increase at 510 nm resulting from the decomposition of H2O2. One unit of enzyme activity is represented by the decomposition of one mmol of H2O2 per minute at 25°C and pH 7.0 under the specified conditions.
- The OxiRed probe method. For this method, POD catalyzes the reaction between OxiRed probe and H2O2 to produce a red fluorescent oxidation product, resorufin. The amount of resorufin is quantified by colorimetric (OD = 570 nm) or fluorometric (Ex/Em = 535/587 nm) approaches. One unit of POD activity is defined as the amount of enzyme that reduces 1.0 mmol of H2O2 per minute at 37 °C.
Glutathione peroxidase (GPx) activity analysis
The GPx activity analysis is based on the fact that GPx can oxidize the reduced glutathione (GSH) to oxidized glutathione (GSSG) at the presence of H2O2. The generated GSSG is recycled to its reduced state, GSH, by glutathione reductase (GR) and NADPH to generate NADP+ (Figure 1). The GPx activity can be measured based on the decrease of NADPH (absorbance at 340 nm). One unit of GPx activity is defined as 1 mmol of NADPH oxidized per minute at 37 °C. Alternatively, the GPx activity can also be quantified by the fluorescence signal of NADP+ using NADP probe (Ex/Em = 420/480 nm).
Lifeasible, as a specialist in plant physiology, provides state-of-the-art methods for antioxidant enzymes analysis. Our experienced project managers will provide you with professional support to ensure the success of your project. Please feel free to contact with Lifeasible for more information.
Reference
- Weydert C. J.; Cullen J. J. Measurement of superoxide dismutase, catalase, and glutathione peroxidase in cultured cells and tissue. Nat Protoc. 2009, 5(1): 51-66.
For research or industrial raw materials, not for personal medical use!
 Figure 1. Functions of three antioxidant enzymes--catalase, superoxide dismutase, and peroxidase (represented by glutathione peroxidase (GPx)) (Weydert and Cullen, 2009).
Figure 1. Functions of three antioxidant enzymes--catalase, superoxide dismutase, and peroxidase (represented by glutathione peroxidase (GPx)) (Weydert and Cullen, 2009).